Tabular Synthetic Data Generation using CTGAN
Checkout mdedit.ai, AI powered Markdown Editor for tech writers
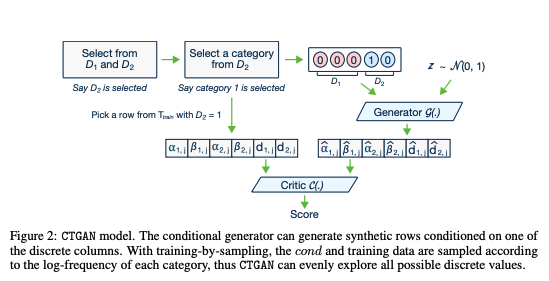
In this post we will talk about generating synthetic data from tabular data using Generative adversarial networks(GANs). We will be using the default implementation of CTGAN [1] model.

Introduction
In the last post on GANs we saw how to generate synthetic data on Synthea dataset. Here’s a link to the post for a refresher:
https://www.maskaravivek.com/post/gan-synthetic-data-generation/
Similar to the last post, we would be working with the Synthea dataset which is publicly available.
https://synthetichealth.github.io/synthea/
In this post, we will be working on the patients.csv file and will only be using continious and categorical fields. We will remove the other fields like name, email ID etc which contains a lot of unique values and will thus will be difficult to learn.
Data Preprocessing
Firstly, download the publicly available synthea dataset and unzip it.